Tag: Phylogenetics
-

Deciding how to construct a phylogenetic tree
For any microbiology project, constructing a phylogenetic tree is an important analysis. A confusing one too — because there are many decisions to make along the way. These decisions can be both technical and hypothesis-driven. How to Build a Dataset for Phylogeny Let’s say someone has isolated and sequenced a few microbial strains. The most…
-

Creating a Publication Quality Phylogeny Using ggtree
A decade ago, circa 2012-2013, I used MEGA5 to infer phylogeny using simple Neighbour-Joining methods, and used the figure generated by MEGA5 to present and publish my results. Later, when I started learning other phylogeny reconstruction methods like Maximum Likelihood (ML) and Bayesian (which does not draw the tree for you), I started to explore…
-
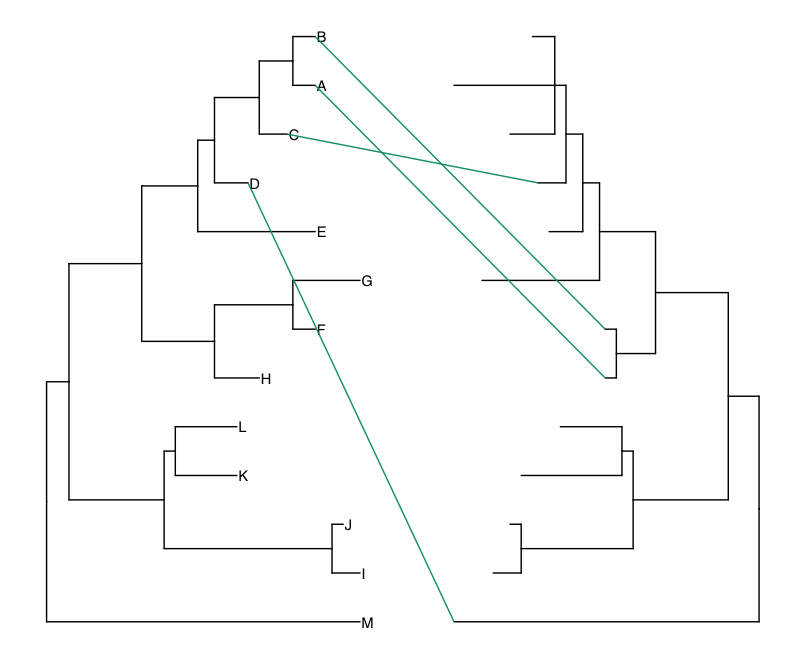
How to make Co-phylogeny plot: easy tanglegram in R
Tanglegrams are co-phylogeny which is a very powerful visualization tool to examine co-evolution. Here is a tutorial on how to make them in R.
